|
x |
x |
|
 |
 |
|
INFECTIOUS
DISEASE |
BACTERIOLOGY |
IMMUNOLOGY |
MYCOLOGY |
PARASITOLOGY |
VIROLOGY |
|
|
BACTERIOLOGY - CHAPTER NINE
GENETIC REGULATORY MECHANISMS
Dr. Gene Mayer
Emeritus Professor
University of South Carolina School of Medicine
|
|
TURKISH |
|
SPANISH |
|
ALBANIAN |
|
|
Let us know what you think
FEEDBACK |
|
SEARCH |

 |
|
Logo image © Jeffrey
Nelson, Rush University, Chicago, Illinois and
The MicrobeLibrary |
|
TEACHING
OBJECTIVES
To discuss the structure
and transcription of bacterial genes
To describe the
molecular mechanisms bacteria use to regulate gene activity
To compare and
contrast inducible and repressible operons
To describe the
molecular mechanisms involved in catabolite repression and attenuation
To discuss the ways
bacteria regulate enzyme activity |
REGULATION OF GENE EXPRESSION
Bacteria do not make all the proteins that they are capable of
making all of the time. Rather, they can adapt to their environment and make
only those gene products that are essential for them to survive in a particular
environment. For example, bacteria do not synthesize the enzymes needed to make tryptophan when there is an abundant supply of tryptophan in the environment.
However, when tryptophan is absent from the environment the enzymes are made.
Similarly, just because a bacterium has a gene for resistance to an antibiotic
does not mean that that gene will be expressed. The resistance gene may only be
expressed when the antibiotic is present in the environment.
Bacteria usually control gene expression by regulating the level of
mRNA transcription. In bacteria, genes with related function are generally located
adjacent to each other and they are regulated coordinately (i.e. when one
is expressed, they all are expressed). Coordinate regulation of clustered genes
is accomplished by regulating the production of a polycistronic mRNA (i.e.
a large mRNA containing the information for several genes). Thus, bacteria are able
to "sense" their environment and express the appropriate set of
genes needed for that environment by regulating transcription of those genes.
|
|
KEY
WORDS
Coordinate gene
expression
Polycistronic m-RNA
Promoter
Operon
Inducible operon
Inducer
Structural gene
Regulatory gene
Repressor
Operator
Negative control
Catabolite repression
CAP protein
Positive control
Repressible operon
Co-repressor
Apo-repressor
Attenuation
Leader region
Feedback inhibition
Epigenetic modification
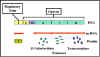 Figure 1 The lactose operon
Figure 1 The lactose operon
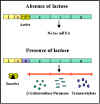 Figure 2 Transcription of lac genes in the presence and absence of
glucose
Figure 2 Transcription of lac genes in the presence and absence of
glucose |
Inducible genes - The operon model
Definition
An inducible gene is a gene that is expressed in the presence of a
substance (an inducer) in the environment. This substance can control the expression of one or
more genes (structural genes) involved in the metabolism of that substance.
For example, lactose induces the expression of the lac genes that are
involved in lactose metabolism. An certain antibiotic may induce
the expression of a gene that leads to resistance to that antibiotic.
Induction is common in metabolic pathways that result in the catabolism of
a substance and the inducer is normally the substrate for the pathway.
Lactose Operon
Structural genes
The lactose operon (figure 1) contains three structural
genes that code for enzymes involved in lactose metabolism.
- The lac z
gene codes for β-galactosidase, an enzyme that breaks down lactose into
glucose and galactose
- The lac y gene codes for a permease,
which is involved in uptake of lactose
- The lac a gene codes for a galactose transacetylase.
These genes are transcribed from a
common promoter into a polycistronic mRNA, which is translated to yield the
three enzymes.
Regulatory gene
The expression of the structural genes is not
only influenced by the presence or absence of the inducer, it is also
controlled by a specific regulatory gene. The regulatory gene may be next to
or far from the genes that are being regulated. The regulatory gene codes
for a specific protein product called a REPRESSOR.
Operator
The repressor acts by binding to a specific region
of the DNA called the operator which is adjacent to the structural genes
being regulated. The structural genes together with the operator region and
the promoter is called an OPERON. However, the binding of the repressor to
the operator is prevented by the inducer and the inducer can also remove
repressor that has already bound to the operator. Thus, in the presence of
the inducer the repressor is inactive and does not bind to the operator,
resulting in transcription of the structural genes. In contrast, in the
absence of inducer the repressor is active and binds to the operator,
resulting in inhibition of transcription of the structural genes. This kind
of control is referred to a NEGATIVE CONTROL since the function of the
regulatory gene product (repressor) is to turn off transcription of the
structural genes.
Inducer
Transcription of the lac genes is influenced by the
presence or absence of an inducer (lactose or other β-galactosides)
(Figure 2).
| e.g |
+ inducer |
expression |
|
- inducer |
no expression |
|
|
CHIME
 Click on image at left to see a rotable molecular structure of the lac
repressor bound to the DNA of the lac operon.
Requires
Netscape and a Chime plug-in. Get Chime here)
Click on image at left to see a rotable molecular structure of the lac
repressor bound to the DNA of the lac operon.
Requires
Netscape and a Chime plug-in. Get Chime here) |
 Figure 3 Catabolite repression
Figure 3 Catabolite repression
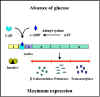 Figure 4 Effect of glucose on expression of proteins encoded by the lac
operon
Figure 4 Effect of glucose on expression of proteins encoded by the lac
operon
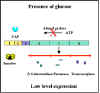 Figure 5 Effect of glucose on expression of proteins encoded by the lac
operon
Figure 5 Effect of glucose on expression of proteins encoded by the lac
operon
|
Catabolite repression (Glucose Effect)
Many inducible operons are not only controlled by their respective
inducers and regulatory genes, but they are also controlled by the level of
glucose in the environment. The ability of glucose to control the expression
of a number of different inducible operons is called CATABOLITE REPRESSION.
This is illustrated in Figure 3.
Catabolite repression is generally seen in those operons which are
involved in the degradation of compounds used as a source of energy. Since
glucose is the preferred energy source in bacteria, the ability of glucose
to regulate the expression of other operons ensures that bacteria will
utilize glucose before any other carbon source as a source of energy.
Mechanism
There is an inverse relationship between glucose levels
and cyclic AMP (cAMP) levels in bacteria. When glucose levels are high cAMP levels are
low and when glucose levels are low cAMP levels are high. This
relationship exists because the transport of glucose into the cell
inhibits the enzyme adenyl cyclase which produces cAMP. In the bacterial
cell cAMP binds to a cAMP binding protein called CAP or CRP. The cAMP-CAP
complex, but not free CAP protein, binds to a site in the promoters of
catabolite repression-sensitive operons. The binding of the complex
results in a more efficient promoter and thus more initiations of
transcriptions from that promoter as illustrated in Figures 4 and 5. Since
the role of the CAP-cAMP complex is to turn on transcription this type of
control is said to be POSITIVE CONTROL. The consequences of this type of
control is that to achieve maximal expression of a catabolite repression
sensitive operon glucose must be absent from the environment and the
inducer of the operon must be present. If both are present, the operon will
not be maximally expressed until glucose is metabolized. Obviously, no
expression of the operon will occur unless the inducer is present.
|
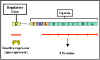 Figure 6 The tryptophan operon
Figure 6 The tryptophan operon
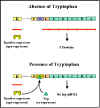 Figure 7 The effect of tryptophan on expression from the tryp operon
Figure 7 The effect of tryptophan on expression from the tryp operon |
Repressible genes - The operon model
Definition
Repressible genes are those in which the presence of a
substance (a co-repressor) in the environment turns off the expression of
those genes (structural genes) involved in the metabolism of that
substance.
e.g., Tryptophan represses the expression of the trp genes.
Repression is common in metabolic pathways that result in the
biosynthesis of a substance and the co-repressor is normally the end product
of the pathway being regulated.
Tryptophan operon
Structural genes
The tryptophan operon
(figure 6) contains five
structural genes that code for enzymes involved in the synthesis of
tryptophan. These genes are transcribed from a common promoter into a
polycistronic mRNA, which is translated to yield the five enzymes.
Regulatory gene
The expression of the structural genes is
not only influenced by the presence or absence of the co-repressor, it is
also controlled by a specific regulatory gene. The regulatory gene may be
next to or far from the genes that are being regulated. The regulatory
gene codes for a specific protein product called an REPRESSOR (sometimes
called an apo-repressor). When the repressor is synthesized it is
inactive. However, it can be activated by complexing with the co-repressor
(i.e. tryptophan).
Operator
The active repressor/co-repressor complex acts by
binding to a specific region of the DNA called the operator which is
adjacent to the structural genes being regulated. The structural genes
together with the operator region and the promoter is called an OPERON.
Thus, in the presence of the co-repressor the repressor is active and
binds to the operator, resulting in repression of transcription of the
structural genes. In contrast, in the absence of co-repressor the
repressor is inactive and does not bind to the operator, resulting in
transcription of the structural genes. This kind of control is referred to
a NEGATIVE CONTROL since the function of the regulatory gene product
(repressor) is to turn off transcription of the structural genes.
Co-repressor
Transcription of the tryptophan genes is
influenced by the presence or absence of a co-repressor (tryptophan)
(Figure 7).
| e.g.
|
+ co-repressor |
no expression |
|
- co-repressor |
expression |
|
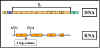 Figure 8 Mechanism of attenuation
Figure 8 Mechanism of attenuation
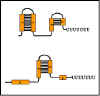 Figure 9 Formation of stem-loops
Figure 9 Formation of stem-loops |
Attenuation
In many repressible operons, transcription that initiates at the promoter
can terminate prematurely in a leader region that precedes the first
structural gene. (i.e. the polymerase terminates transcription before
it gets to the first gene in the operon). This phenomenon is called
ATTENUATION; the premature termination of transcription. Although
attenuation is seen in a number of operons, the mechanism is best understood
in those repressible operons involved in amino acid biosynthesis. In these
instances attenuation is regulated by the availability of the cognate
aminoacylated t-RNA.
Mechanism (See Figure 8 )
When transcription is initiated at the promoter, it actually starts
before the first structural gene and a leader transcript is made. This
leader region contains a start and a stop signal for protein synthesis.
Since bacteria do not have a nuclear membrane, transcription and
translation can occur simultaneously. Thus, a short peptide can be made
while the RNA polymerase is transcribing the leader region. The test
peptide contains several tryptophan residues in the middle of the peptide.
Thus, if there is a sufficient amount of tryptophanyl-t-RNA to translate
that test peptide, the entire peptide will be made and the ribosome will
reach the stop signal. If, on the other hand, there is not enough
tryptophanyl-t-RNA to translate the peptide, the ribosome will be arrested
at the two tryptophan codons before it gets to the stop signal.
The sequence in the leader m-RNA contains four regions, which have
complementary sequences (Figure 9). Thus, several different secondary stem
and loop structures can be formed. Region 1 can only form base pairs with
region 2; region 2 can form base pairs with either region 1 or 3; region 3
can form base pairs with region 2 or 4; and region 4 can only form base
pairs with region 3. Thus three possible stem/loop structures can be
formed in the RNA.
region 1:region 2
region 2:region 3
region 3:region 4
One of the possible structures (region 3 base pairing with region 4)
generates a signal for RNA polymerase to terminate transcription (i.e.
to attenuate transcription). However, the formation of one stem and loop
structure can preclude the formation of others. If region 2 forms base
pairs with region 1 it is not available to base pair with region 3.
Similarly if region 3 forms base pairs with region 2 it is not available
to base pair with region 4.
The ability of the ribosomes to translate the test peptide will affect
the formation of the various stem and loop structures Figure 10. If the
ribosome reaches the stop signal for translation it will be covering up
region 2 and thus region 2 will not available for forming base pairs with
other regions. This allows the generation of the transcription termination
signal because region 3 will be available to pair with region 4. Thus,
when there is enough tryptophanyl-t-RNA to translate the test peptide
attenuation will occur and the structural genes will not be transcribed.
In contrast, when there is an insufficient amount of tryptophanyl-t-RNA to
translate the test peptide no attenuation will occur. This is because the
ribosome will stop at the two tryptophan codons in region 1, thereby
allowing region 2 to base pair with region 3 and preventing the formation
of the attenuation signal (i.e. region 3 base paired with region
4). Thus, the structural genes will be transcribed.
|
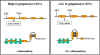 Figure 10 Mechanism of attenuation
Figure 10 Mechanism of attenuation |
| |
REGULATION OF ENZYME ACTIVITY
Bacteria also have ways of regulating the activities of their enzymes.
Feedback Inhibition
The activity of bacterial enzymes is
often subject to feedback inhibition. Usually it is the end product of a
pathway that is the inhibitor and the first enzyme in the pathway is the step
that is regulated.
Epigenetic Modification
The activities of bacterial enzymes can
also be regulated by covalent modifications of enzymes. Such modifications are
called EPIGENETIC MODIFICATIONS.
e.g. Adenylation of glutamine synthetase
Phosphorylation of glycogen synthetase
Usually these modifications are reversible so that the activities of the
enzymes can be turned on and off.
|
|
|
 Return to the Bacteriology Section
of Microbiology and Immunology On-line Return to the Bacteriology Section
of Microbiology and Immunology On-line
This page last changed on
Wednesday, March 02, 2016
Page maintained by
Richard Hunt
|

 Figure 3 Catabolite repression
Figure 3 Catabolite repression
 Figure 6 The tryptophan operon
Figure 6 The tryptophan operon
 Figure 10 Mechanism of attenuation
Figure 10 Mechanism of attenuation






